Recognizing synapses in the zfish dataset
To identify synapses in zfish cells, look for 2 things:
- A short stretch of border between 2 cells, that is darker and thicker than the rest of the border.
- An area of increased neurotransmitter vesicle density on one side of the dark border.
When you've identified a synapse, it should be easy to tell which side of the border is the axon and which is the dendrite. The area of higher vesicle density marks the presynaptic neurite, or axon. The other side is the postsynaptic neurite, or dendrite.
Note: there is such a thing as postsynaptic density, but I'm not sure if that is visible in zfish slides. Ironically, the presynaptic side is visibly denser than the postsynaptic side due to the neurotransmitter vesicles.
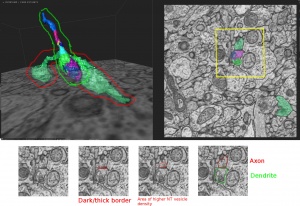
Below is an example from cube #2134873, cell 77327.
The axon is circled red and the dendrite circled green. The yellow square roughly indicates the borders of the cropped images in the bottom row. Each of those images is the same image with different annotations drawn on them (except for the first, which allows you to see the original greyscale micrograph with no obstructing highlights/markings). The bottom row of images, from left to right, show:
- Original greyscale image
- Dark/thick border (circled red), which along with NT vesicles in #3, indicates a possible synapse
- Area of higher neurotransmitter vesicle density (circled red), which indicates the axon
- Axon (outlined in red) vs dendrite (outlined in green)
Many more examples will appear here over time.