E2198
yeWire se basa en las imágenes que se adquirieron en el Instituto Max Planck para la Investigación Médica en Heidelberg, Alemania. Este conjunto de datos, conocido como E2198, fue la base de la especificidad de cableado en el circuito de selectividad de dirección de la retina. Cite error: Invalid <ref> tag;
invalid names, e.g. too many La fuente retina se obtuvo de un adulto natural adaptado a la oscuridad (C57BL/6) ratón.
Los investigadores midieron tanto la actividad neural como la conectividad en la misma retina mediante la aplicación de dos métodos de imagen uno tras otro: microscopía de dos fotones (2P) y microscopía electrónica de barrido de cara en serie (SBFSEM).
Hoy en día es común estimar la tasa de espina neural mediante la imagen de la concentración de calcio intracelular. Esta estimación tiene sentido porque a cada pico le sigue una entrada de calcio a través de canales de calcio sensibles al voltaje. La concentración de calcio se puede visualizar mediante 2P después de cargar las células con un tinte (por ejemplo, Oregon Green BAPTA) que cambia su fluorescencia en función de la concentración de calcio.
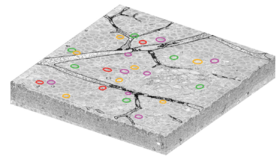
De esta manera, Briggman et al. imaginó las respuestas de las células ganglionares con 2P mientras aplicaba estímulos visuales a una retina de ratón adulto. Las manchas blancas en la imagen de la derecha son las ganglio cuerpos celulares, y las ramas oscuras son vasos sanguíneos. Los círculos de colores marcan las ubicaciones de Célula de Ganglio Selectivo de Dirección de Encendido Apagado. Estas células ganglionares selectivas en dirección (DSGC) se dividen en cuatro tipos, cada una de las cuales es selectiva para una de las cuatro direcciones cardinales, y están marcadas por cuatro colores (magenta, verde, rojo y amarillo).
Immediately after 2P imaging, the retina was fixed, stained, and embedded in a hard plastic resin. An unconventional stain was used to mark the boundaries between neurons while leaving intracellular organelles unstained. SBFSEM was used to image a volume of size 350×300×60 µm3 (left) with voxel resolution 16.5×16.5×23 nm3. Since the same blood vessels were visible in both the 2P and SBFSEM images, the researchers were able to find the same DSGCs in both images. These are marked by colored circles on the SBFSEM image just as in the 2P image above.
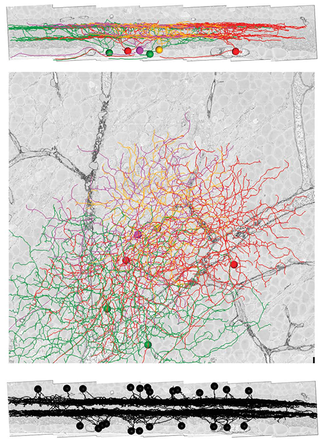
In Panel a of the figure on the right, the gray scale image shows a cross section of the SBFSEM volume that is perpendicular to the retina. You can think of this as a cut through a sandwich. The filling is the inner plexiform layer (IPL), the top piece of bread the inner nuclear layer (INL) and the bottom piece of bread the ganglion cell layer (GCL). The IPL contains synaptic connections between the axons of bipolar cells, and the dendrites of amacrine and ganglion cells.
The colored objects in panel a are six DSGCs reconstructed by the researchers. The circles are representations of the cell bodies, and the lines are "skeletons" of the dendrites. Each DSGC is said to be "bistratified," which means that its dendrites branch out in two sublayers ("strata") of the IPL. The total number of strata in the IPL is estimated to be around ten. A view of the bottom of the sandwich (panel b) shows the branching of the DSGC dendrites.
The researchers also reconstructed 24 starburst amacrine cells (SACs), which are shown in Panel c. These cells fall in two classes: ON cells (n=11) stratifying in one sublayer of the IPL, and OFF cells (n=13) stratifying in another sublayer. Comparison with panel a shows that SAC dendrites co-stratify with DSGC dendrites, suggesting that these two cell types could be connected by synapses.
As shown in the images below, SAC dendrites and DSGC dendrites indeed contact each other. However, not all points of contact are synapses. The researchers conjectured that "varicose" contacts, at which one dendrite seems to grasp or embrace another, were actual synapses, while "incidental" contacts were not synapses.
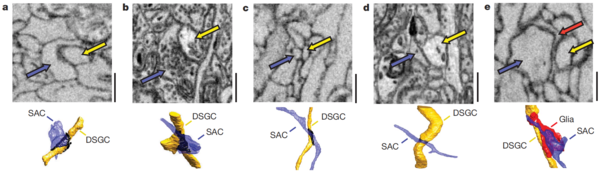
From this analysis, the researchers derived a rule governing DSGC-SAC connectivity. All SAC dendrites and DSGCs respond selectively to ("prefer") motion in some direction. The researchers found that DSGCs tend to receive more synapses from SAC dendrites with the opposite preferred direction, which is evidence for specificity of connections. In principle, a DSGC could indiscriminately receive synapses from all neighboring SAC dendrites, which generally encompass a diversity of preferred directions, but this is not the case. The rule of connectivity suggests that a DSGC "inherits" its direction selectivity from its inputs, since SACs make inhibitory (sign-inverting) synapses onto DSGCs.
This study was based on reconstructions of just 30 neurons in E2198. Clearly there is much more to be learned from this dataset, which contains thousands of neurons.
Training Data
You can access the original convolutional neural network training data here: